Simons Machine Learning Center
at the New York Structural Biology Center
The Simons Machine Learning Center (SMLC) is a Simons Foundation funded initiative and is part of the Simons Electron Microscopy Center at the New York Structural Biology Center.
We are building a team of machine learning, computer vision, and computational biology experts to push the boundaries of machine learning and structural biology. Machine learning is transforming protein and structural biology, but new methods are needed to fully integrate machine learning into the structural biology workflow and to incorporate biophysical knowledge into our algorithms. Our mission is to develop machine learning algorithms and software for better understanding and predicting the structures of biological molecules and for linking those structures with functional properties. Our interests span algorithms for accelerating structure determination and image analysis to methods for modeling protein sequence, structure, function, and beyond. As part of NYSBC, we have unprecedented access to state-of-the-art instruments, technologists, and structural biologists.
Learn more about our research.
Resources:
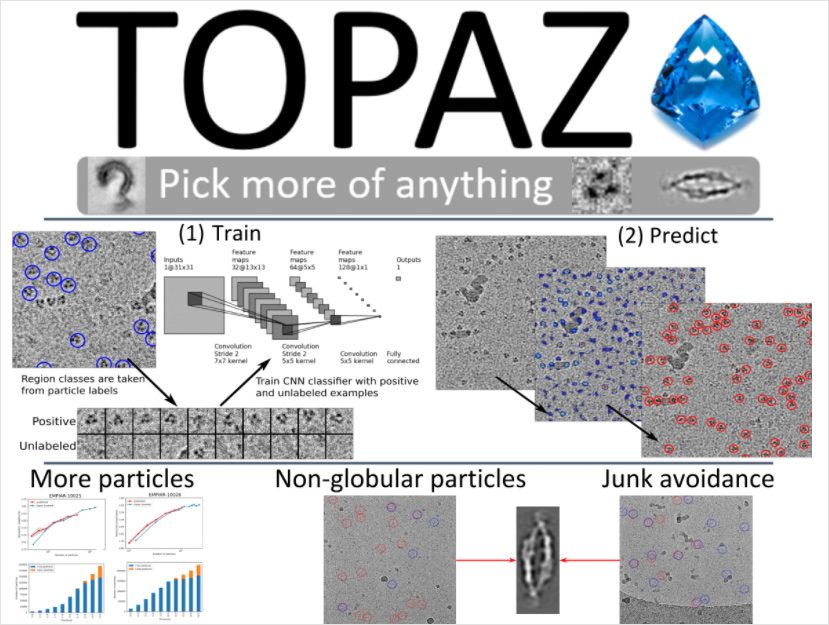
Topaz and Topaz-Denoise
Topaz is a software package for particle picking in cryo-electron micrographs using convolutional neural networks and positive-unlabeled learning. Topaz is free and open source and contains pretrained general purpose particle picking models, a stand alone GUI, and cryoSPARC, RELION, and Scipion integration. Topaz now also includes neural networks for micrograph and tomogram denoising with Topaz-Denoise.
Spatial-VAE (continuous classification)
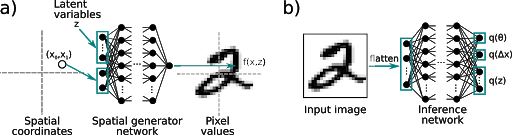
Spatial-VAE is a method for learning continuous latent variable models of images that are robust to uninteresting pose transformations (i.e. rotation and translation). This method is useful for continuous 2d classification of flexible proteins in cryoEM images and has been further developed into a complete 3d reconstruction solution in cryoDRGN by Ellen Zhong at MIT.
